OpenAI just released a set two tools to use for tasks related to biochemistry and medicine. They will make your life much easier if you are working in this field.
The first one is a specialized AI model that is great at exploring problems related to life sciences. This one is currently only available to research institutions working with OpenAI. I will let you know here in the channel as soon as it is widely available. For now, we will go over the benchmarks briefly and talk about what makes it different.
The second one is a tool you can start using today. It’s a specialized, open-source plugin that you can use with any AI agent to help you with tasks in the life sciences. I’m going to show you how to set it up with OpenAI models, Claude Code, and open source models for free.
Let’s get right into it.
GPT-Rosalind#
First, let’s quickly go over GPT-Rosalind.
This is a fine-tuned version of OpenAI’s latest model, GPT-5.4, which is a foundation model already capable of solving complex problems across multiple fields. With the new Rosalind version, it now has increased performance across a few benchmarks relevant to scientific research.
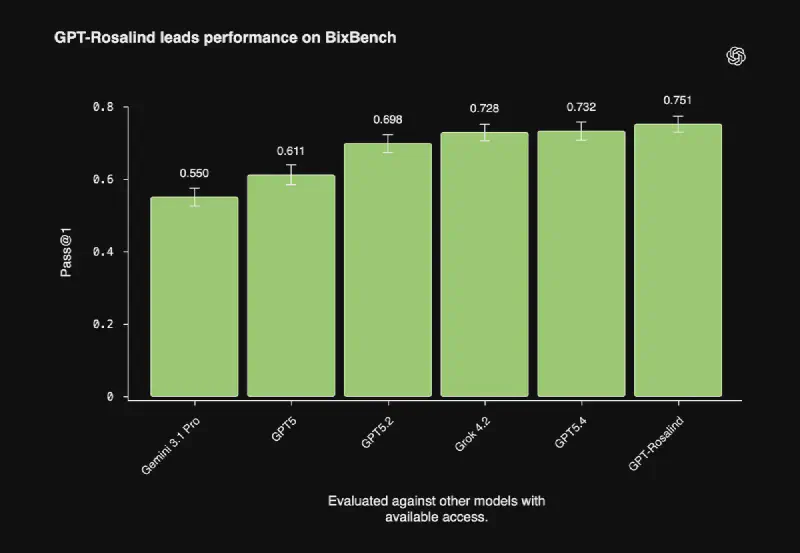
BixBench, the benchmark showed in the plot above, measures how well a model performs on a variety of real-world bioinformatics and data analysis tasks. As you can see in the plot, it performs better on this benchmark than the previously leading model, which is GPT-5.4.
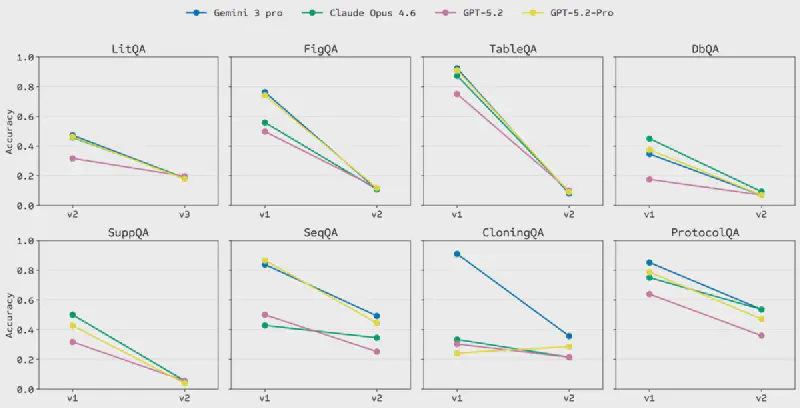
On LABBench2, a benchmark measuring performance on a range of research tasks such as literature retrieval, database access, sequence manipulation and protocol design, GPT-Rosalind outperforms GPT-5.4 on 6 out of 11 tasks.
To request access to GPT-Rosalind, you can fill out the form on OpenAI’s website. I will let you know here on the channel as soon as it is widely available.
Life Science Research Plugin#
Now this is where the fun starts. This is a plugin that you can use with any AI coding agent. It is completely open source (so you know exactly what your AI agent is consuming), and it is free.
With it, you can ask Codex (the AI coding agent from OpenAI), Claude Code or any open model, questions like:
- “Summarize what genes most strongly link triglycerides and NAFLD. Focus on the strongest genetics, expression, and pathway evidence, and call out any caveats.”
- “Explain why PCSK9 is a compelling drug target. Include protein function, disease relevance, known drugs, and any important caveats.”
I am not an expert in any of these topics, but I asked Claude Code with Sonnet 4.5, Codex with GPT-5.4, and MiniMax M2.7 the first question of the above examples, and asked them to produce a PDF report with their findings. All of them used the plugin to explore research databases and produce a detailed report.
Here are the results:
- Claude Code:
- Codex:
- MiniMax:
I’m not an expert in any of these topics, so feel free to judge yourself which model produced the best report.
How to install the plugin#
OpenAI Codex#
If you are paying for a premium Codex plan, you can just open the Codex app and click on Plugins, search for Life Science Research, and click on Install.
Then, Codex will be able to identify when it needs to use the plugin. Or you can explicitly call the plugin by clicking on the bottom “+” sign.
Claude Code and Open Models#
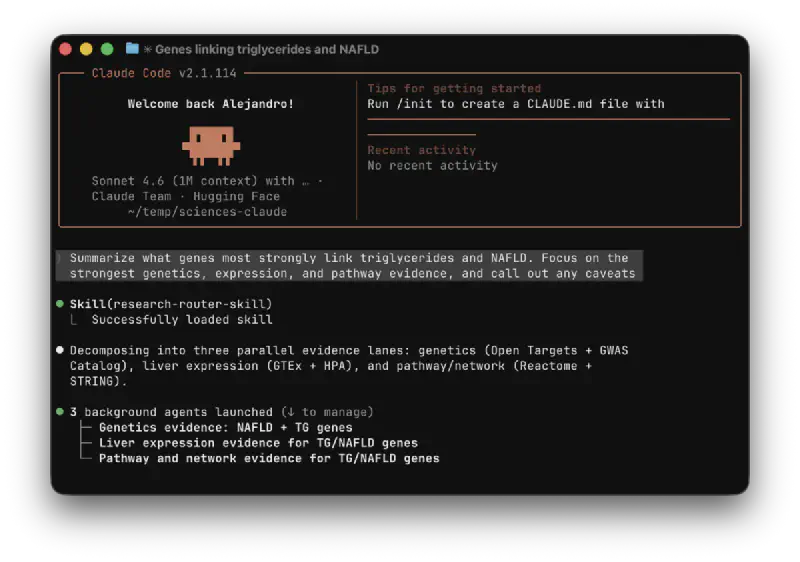
If you have a premium Claude account, you can install the skills from this plugin with skills.sh.
To do this, open your command line where you will be working with life science projects, and run:
npx skills add https://github.com/openai/plugins/tree/main/plugins/life-science-research
Follow the instructions and make sure to select all the skills in the plugin by clicking on a when prompted to select the skills you want.
If you want to use this with Claude Code, be sure to select Claude Code when prompted about the agent that you will be using.
Once that’s done, feel free to run it with your agent.
Using with Open Models#
If you run out of tokens from one of the above two agents, you can use open models, which are much more affordable and are often as good or better than the agents from OpenAI or Anthropic. I personally use open models as my daily agents.
We will use OpenCode as the agent harness and we will plug open models from Hugging Face.
Install OpenCode#
If you want to use this with open models, my recommendation is to use an open source agent like OpenCode. You can plug it to open source models, which are more affordable, and also about as powerful as OpenAI’s or Anthropic’s models.
To install OpenCode, just run the following command in your terminal:
curl -fsSL https://opencode.ai/install | bash
And when you open it, select the models that you want to use by running:
/connect
Use Open Models from Hugging Face#
I recommend that you use Hugging Face’s Inference Providers to connect to many open source models without any markup price. Just create your account at hf.co and create a token at https://huggingface.co/settings/tokens.
Then you can add that token to the OpenCode assistant and select any model you want. I recommend that you use either Kimi K2.5 or Minimax M2.7 or M2.5.
Research Plugin with OpenCode#
If you followed the steps from the Claude Code section, you should also have the OpenCode skills installed. So you can now just query OpenCode and use it when you run out of tokens in Codex or Claude Code.
Conclusion#
I hope this was a useful tutorial on running life sciences research plugin on different AI agents. Feel free to let me know if you have any questions.

